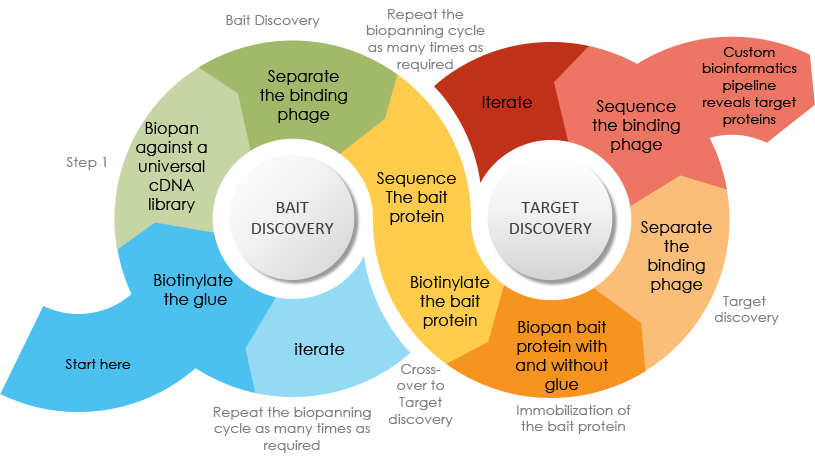
Our technology allows a highly parallel approach to finding potential targets of drug candidates, new drug modalities, and enables more informed decisions in drug discovery.
Using Mercurious®, the Company believes scientists can discover the true cellular target(s) for bioactive compounds (small molecule drugs, drug leads and biologics) with high specificity and high affinity more rapidly than by conventional biochemical approaches. The Company’s technology is complementarity to other target identification technologies such as affinity capture mass spectrometry, CRISPR-screens and yeast 3-hybrid but has some unique advantages due to the iterative nature of the process. The Mercurious platform can also be used to discover molecular glues and the targets for molecular glues and the Company is actively pursuing this space for the development of first-in-class new modalities.
Our lead program (MB2052) in oncology binds the “undruggable” KIX domain or EP300 which is progressing from discovery to in vitro validation. We have several other programs targeting molecular glues for hard-to-drug transcription factors and co-regulators that are in the discovery phase.
About 60% of proteins are common to all tissue types and only 3% are specific to one cell type – and almost all of these (94%) are low abundance proteins. Unfortunately, these are exactly the proteins we need to identify for drug development. Our Mercurious phage display platform physically links protein domains with their gene to render even the lowest abundance proteins detectable through iterative affinity purification and PCR-like amplification. The Company’s technology can be used to identify all the protein targets of drugs, drug leads, screening hits, natural products and biologics that display promising pharmacological activities that have unknown mechanisms of action, identify possible off-targets that could be related to side effects or allow drug repositioning. We do this through the innovative combination of chemical synthesis and phage biopanning to identify the most avid drug-protein interactions involved. Our approach provides a relatively unbiased method that is complementary of forward proteomics approaches (e.g. affinity capture mass spectrometry, DARTS and CETSA) and genetics methods (e.g. CRISPR-Cas9/sgRNA and shRNA) methods to reveal fundamental insights into the mechanisms of action of biologically active molecules.
Our technology allows a highly parallel approach to finding potential targets of drug candidates and enables fail-fast and fail cheap decisions in drug discovery. By extracting mRNA of a cell or tissue with temporal and spatial control, coding and non-coding transcriptomics information can be packaged as cDNA libraries and displayed on viral (e.g. phage) particle surfaces to generate diverse but ordered collections of functional protein libraries for screening. Bioactive compounds can be iteratively screened against these protein libraries more rapidly than conventional biochemical approaches and generates unique binding matrices for better assessment of targeting affinity and specificity. This approach works for small molecular, biologics and molecular glues and the Company is pursuing it own molecular glue discovery programs using the Mercurious Platform to identify new bait and target proteins from putative molecular glues.

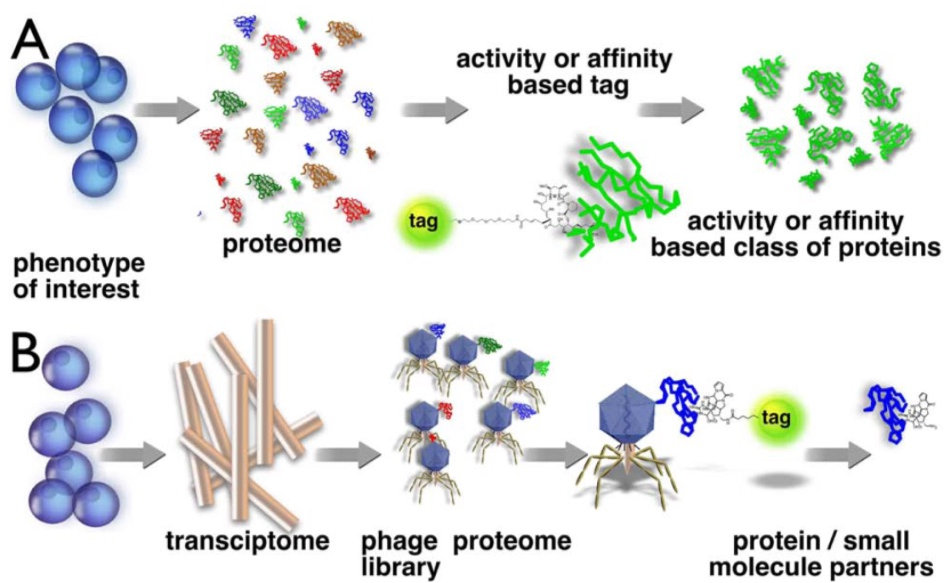
A) Forward chemical proteomics starts with a a proteome interrogated with a tagged drug, pulling down activity of affinity associated proteins and putatively identify the interacting proteins based on the molecular weight of trytpic peptides,
B) Reverse Chemical Proteomics starts with the transcriptome (from mRNA) displayed on the surface of a vector to achieve iterative protein/peptide purification of the most avid binding partner but read out on the DNA level